Temperature minimization
Density-based topology optimization to minimize average temperature.
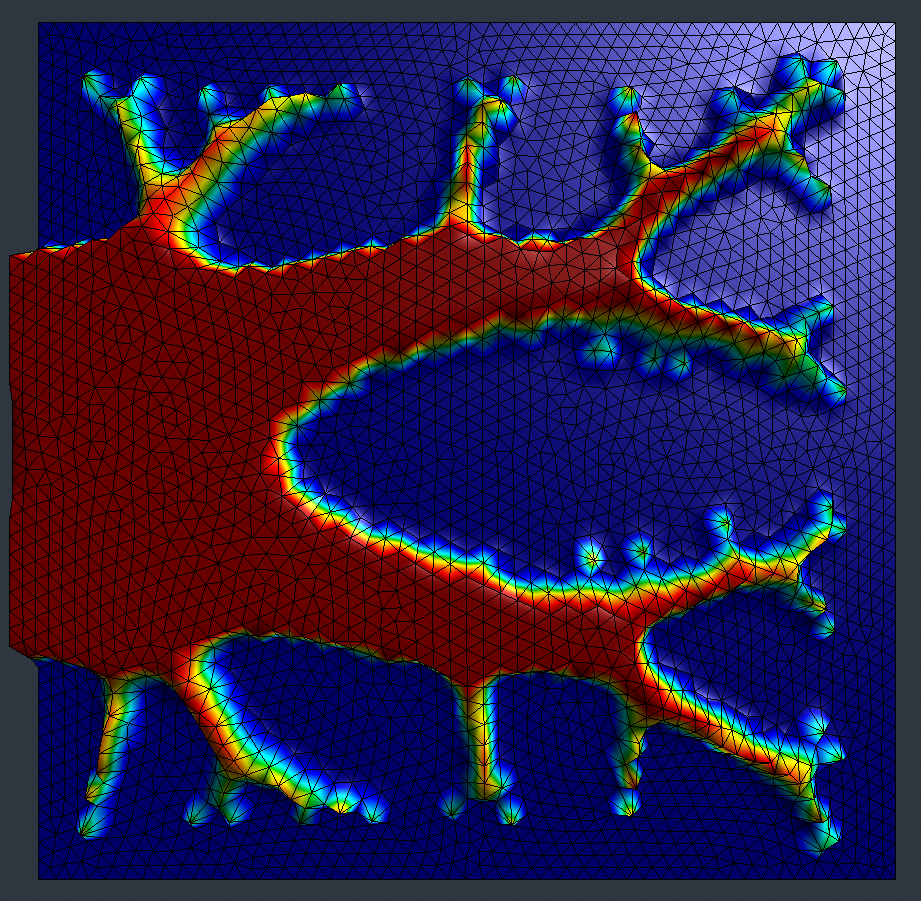
Introduction
This example demonstrates density-based topology optimization using the SIMP (Solid Isotropic Material with Penalization) method. The goal is to find an optimal material distribution that minimizes the average temperature in a domain heated by a uniform source.
The optimization problem reads:
subject to the state equation (a Poisson problem with SIMP-interpolated conductivity):
where the conductivity is interpolated as .
Setup
Parameters
static constexpr int Gamma0 = 1, GammaD = 2; static constexpr double ell = 4; // Lagrange multiplier static constexpr double mu = 0.2; // Step size static constexpr double gmin = 0.0001; // Minimum conductivity static constexpr double gmax = 1; // Maximum conductivity static constexpr double alpha = 0.05; // Regularization parameter static constexpr size_t maxIterations = 5000;
Mesh and Initial Density
The mesh is loaded and optimized with MMG::
MMG::Mesh Omega; Omega.load(meshFile, IO::FileFormat::MFEM); MMG::Optimizer().setHMax(0.005).setHMin(0.001).optimize(Omega); P1 vh(Omega); P1 ph(Omega); GridFunction gamma(ph); gamma = 0.9; // Initial density: nearly full material
Optimization Loop
Each iteration consists of three PDE solves:
Step 1 — State Equation
The temperature field is the solution of a Poisson problem with SIMP-interpolated conductivity :
TrialFunction u(vh); TestFunction v(vh); Problem poisson(u, v); poisson = Integral((gmin + (gmax - gmin) * Pow(gamma, 3)) * Grad(u), Grad(v)) - Integral(f * v) + DirichletBC(u, RealFunction(0.0)).on(GammaD); Solver::SparseLU(poisson).solve();
Step 2 — Adjoint Equation
The adjoint problem is similar to the state equation but with a different right-hand side arising from the derivative of the objective:
TrialFunction p(vh); TestFunction q(vh); Problem adjoint(p, q); adjoint = Integral((gmin + (gmax - gmin) * Pow(gamma, 3)) * Grad(p), Grad(q)) + Integral(RealFunction(1.0 / vol), q) + DirichletBC(p, RealFunction(0.0)).on(GammaD); Solver::SparseLU(adjoint).solve();
Step 3 — Gradient and Update
The topological gradient is smoothed via a Hilbert extension-regularization solve. The density is then updated and projected to :
TrialFunction g(vh); TestFunction w(vh); Problem hilbert(g, w); hilbert = Integral(alpha * Grad(g), Grad(w)) + Integral(g, w) - Integral( ell + 3 * (gmax - gmin) * Pow(gamma, 2) Dot(Grad(u.getSolution()), Grad(p.getSolution())), w) + DirichletBC(g, RealFunction(0.0)).on(GammaD); Solver::SparseLU(hilbert).solve(); GridFunction step(ph); step = mu * g.getSolution(); gamma -= step; gamma = Min(1.0, Max(0.0, gamma));
Full Source Code
The complete source code is:
See Also
- Poisson equation — The underlying PDE
- Shape optimization — Boundary-variation approach
- Gallery